Concept
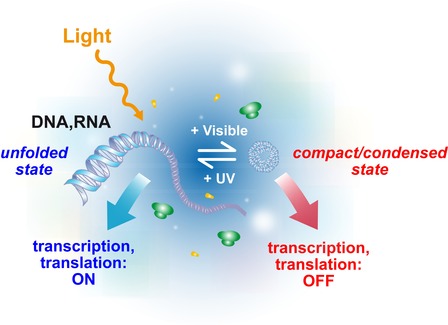
Light has been recognized as an ideal external trigger to address gene expression as it offers unique advantages: high spatio-temporal resolution of the excitation, tunability of the intensity, low perturbation of the biochemical environment, biocompatibility, and high potentiality for biotechnological applications. Various strategies have been proposed in the past few years to control gene expression systems by light, such as photocaged molecules, DNA modification with photoactivable groups, or light-switchable gene promoter systems. All these strategies are based on a sequence-dependent regulation and thus have to be adapted for each particular transcription/translation system. In contrast, we have recently described a novel method of photocontrol of both transcription and translation that is fully reversible, dynamic and potentially applicable to any system of gene expression of interest. In this method, photo-regulated conformation of DNA or mRNA is used as a trigger of transcription and translation reactions (see figure).
Video
Photocontrol of DNA compaction Real-time fluorescence microscopy of T4 DNA (1 µg/mL) dyed with YOYO-1 (100 nM) in transcription conditions (E. coli RNA polymerase 0.02 U/μL, tris-HCl pH 7.5 40 mM, KCl 150 mM, MgCl2 10 mM, NTPs 0.5 mM, T = 37 °C) for different AzoTAB concentrations and UV illumination conditions. Without UV, all DNA moleculeshave an elongated coil conformation ([AzoTAB] = 0 mM and 1.3 mM). DNA molecules fold into a compact state when a sufficient amount of AzoTAB is added ([AzoTAB] = 2 mM). When UV is applied on this sample, all initially compact DNA molecules unfold and recover the elongated coil conformation.
Applications
We are applying this method for the photocontrol of the synthesis of various functional proteins. Following this way we can place under photocontrol the functions provided by the target protein. For instance, by applying this strategy on a beta-lactamase enzyme, we have been able to control with light the conversion of beta-lactams into beta-amino acids. Moreover, we have shown that our method, which is by nature sequence-independent, can be combined with sequence-specific silencing by small RNAs. Following this way, it is also possible to selectively address a target gene using light.
Key-publications
Sequence-independent and reversible photocontrol of transcription/expression systems using a photosensitive nucleic acid binder
A. Estévez-Torres, C. Crozatier, A. Diguet, T. Hara, H. Saito, K. Yoshikawa, D. Baigl, Proc. Natl. Acad. Sci. USA 2009, 106, 12219 - doi : 10.1073/pnas.0904382106 -
The first demonstration of a reversible photocontrol of gene expression based on the use of light-induced nucleic acid conformational changes and Damien's favorite paper
Light-Regulated mRNA Condensation by a Photosensitive Surfactant Works as a Series Photoswitch of Translation Activity in the Presence of Small RNAs
S. Rudiuk, H. Saito, T. Hara, T. Inoue, K. Yoshikawa, D. Baigl*, Biomacromolecules 2011, 12, 3945 - doi : 10.1021/bm200962s -
Where we show that our photocontrol method is compatible with microRNA silencing strategy thus enabling a photocontrol of gene expression with gene selectivity
Sequence-independent and reversible photocontrol of transcription/expression systems using a photosensitive nucleic acid binder
A. Estévez-Torres, C. Crozatier, A. Diguet, T. Hara, H. Saito, K. Yoshikawa, D. Baigl, Proc. Natl. Acad. Sci. USA 2009, 106, 12219 - doi : 10.1073/pnas.0904382106 -
The first demonstration of a reversible photocontrol of gene expression based on the use of light-induced nucleic acid conformational changes and Damien's favorite paper
Light-Regulated mRNA Condensation by a Photosensitive Surfactant Works as a Series Photoswitch of Translation Activity in the Presence of Small RNAs
S. Rudiuk, H. Saito, T. Hara, T. Inoue, K. Yoshikawa, D. Baigl*, Biomacromolecules 2011, 12, 3945 - doi : 10.1021/bm200962s -
Where we show that our photocontrol method is compatible with microRNA silencing strategy thus enabling a photocontrol of gene expression with gene selectivity
Modification-free photocontrol of β-lactam conversion with spatio-temporal resolution
A. Venancio-Marques, Y.-J. Liu, A. Diguet, T. di Maio, A. Gautier, and D. Baigl*, ACS Synth. Biol. 2012, 1, 526–531 - doi : 10.1021/sb300010a - ![]()
Reversible photocontrol of beta-lactam conversion via the photocontrol of beta-lactamase synthesis