Reference: Soft Matter. 2011, 7, pp 61–63.
Title: Dendrimer-induced DNA bending.
Authors: Chun-Yu Chen, Chun-Jen Su, Shu-Fen Peng, Hsin-Lung Chen and Hsing-Wen Sung
Review by: Sergii Rudiuk
Title: Dendrimer-induced DNA bending.
Authors: Chun-Yu Chen, Chun-Jen Su, Shu-Fen Peng, Hsin-Lung Chen and Hsing-Wen Sung
Review by: Sergii Rudiuk
The authors studied structures of DNA-dendrimers complexes (dendriplexes), at different dendrimer charge density.
Experimental system.
Fourth generation polyamidoamine (PAMAM) dendrimer was used for this study. Dendrimers with different degrees of protonation (dp) were prepared by adding different amounts of HCl. The dendriplexe precipitates were prepared by mixing calf thymus DNA and protonated dendrimer solution at N/P ratio fixed at 6/1. SAXS measurements were then performed from aqueous suspensions of dendriplexes.
Results
Dendrimer charge density was expressed in degree of protonation (dp), where: dp = 1 corresponds to all amines protonated, dp = 0 – no protonation, and dp = 0.5 – only surface amines protonated (no interior tertiary amine groups protonated).
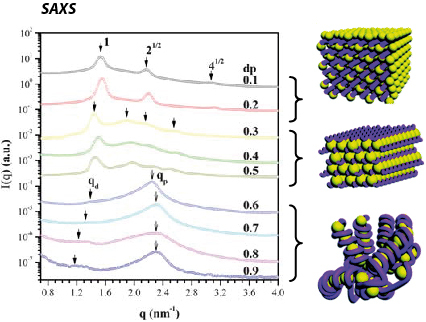
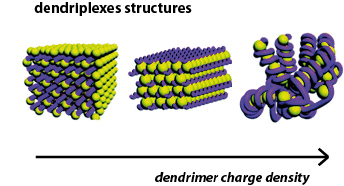
3 distinct nanostructures, characterized by different degrees of DNA bending were determined depending on dendrimer degree of protonation:
- At low dp (0.1 and 0.2), dendriplexes have square columnar phase
- At intermediate dp (0.3-0.5) DNA is packed predominantly in a hexagonal lattice.
- At high dp (≥ 0.6) a chromatin-like fiber was detected having beads-on-string structure. Calculations showed that DNA was wrapped around a dendrimer molecule by ca. 1.4 turns. Here, the strong electrostatic attraction outweighs the DNA bending energy. It has been also shown that DNA penetrated into the dendrimer while wrapping around it, and taht the length of linker DNA between dendrimer "beads" increased with increasing the dendrimer charge density.
Experimental system.
Fourth generation polyamidoamine (PAMAM) dendrimer was used for this study. Dendrimers with different degrees of protonation (dp) were prepared by adding different amounts of HCl. The dendriplexe precipitates were prepared by mixing calf thymus DNA and protonated dendrimer solution at N/P ratio fixed at 6/1. SAXS measurements were then performed from aqueous suspensions of dendriplexes.
Results
Dendrimer charge density was expressed in degree of protonation (dp), where: dp = 1 corresponds to all amines protonated, dp = 0 – no protonation, and dp = 0.5 – only surface amines protonated (no interior tertiary amine groups protonated).
3 distinct nanostructures, characterized by different degrees of DNA bending were determined depending on dendrimer degree of protonation:
- At low dp (0.1 and 0.2), dendriplexes have square columnar phase
- At intermediate dp (0.3-0.5) DNA is packed predominantly in a hexagonal lattice.
- At high dp (≥ 0.6) a chromatin-like fiber was detected having beads-on-string structure. Calculations showed that DNA was wrapped around a dendrimer molecule by ca. 1.4 turns. Here, the strong electrostatic attraction outweighs the DNA bending energy. It has been also shown that DNA penetrated into the dendrimer while wrapping around it, and taht the length of linker DNA between dendrimer "beads" increased with increasing the dendrimer charge density.